Our research group focuses on developing computational approaches to analyze high-throughput drug and genetic screens, along with multi-omics datasets, to study the fundamental regulatory mechanisms underlying cancer cell response to treatments. Working closely with experimental and clinical groups, we develop integrative approaches to address one of the most pressing challenges in cancer: drug resistance.
We aim to develop the next generation of multidisciplinary engineers who can bridge the fields of computer science, cell biology, and biomedicine.
Machine Learning | Multi-omics | Functional Genomics | Drug resistance | Cancer
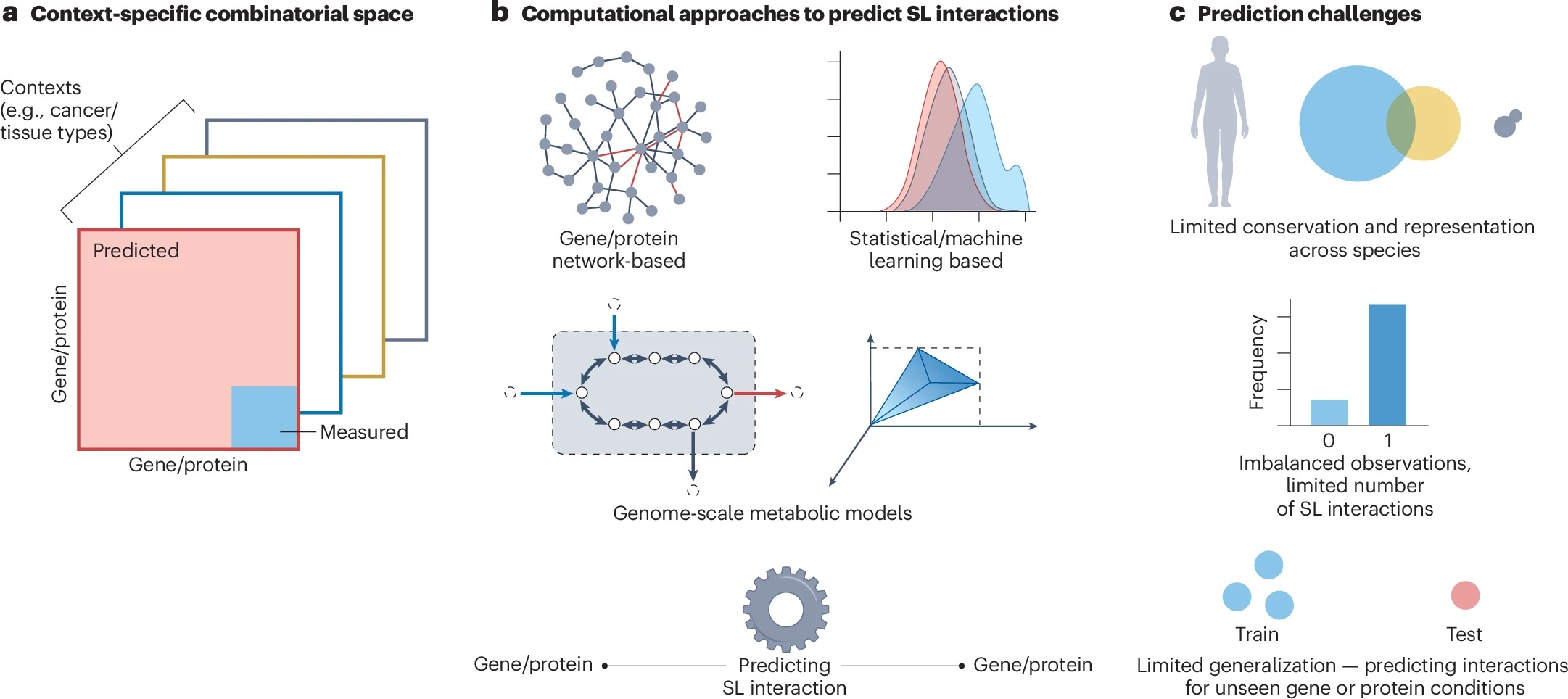
Synthetic lethality, first proposed more than two decades ago, has long held immense promise for...
[Read More]
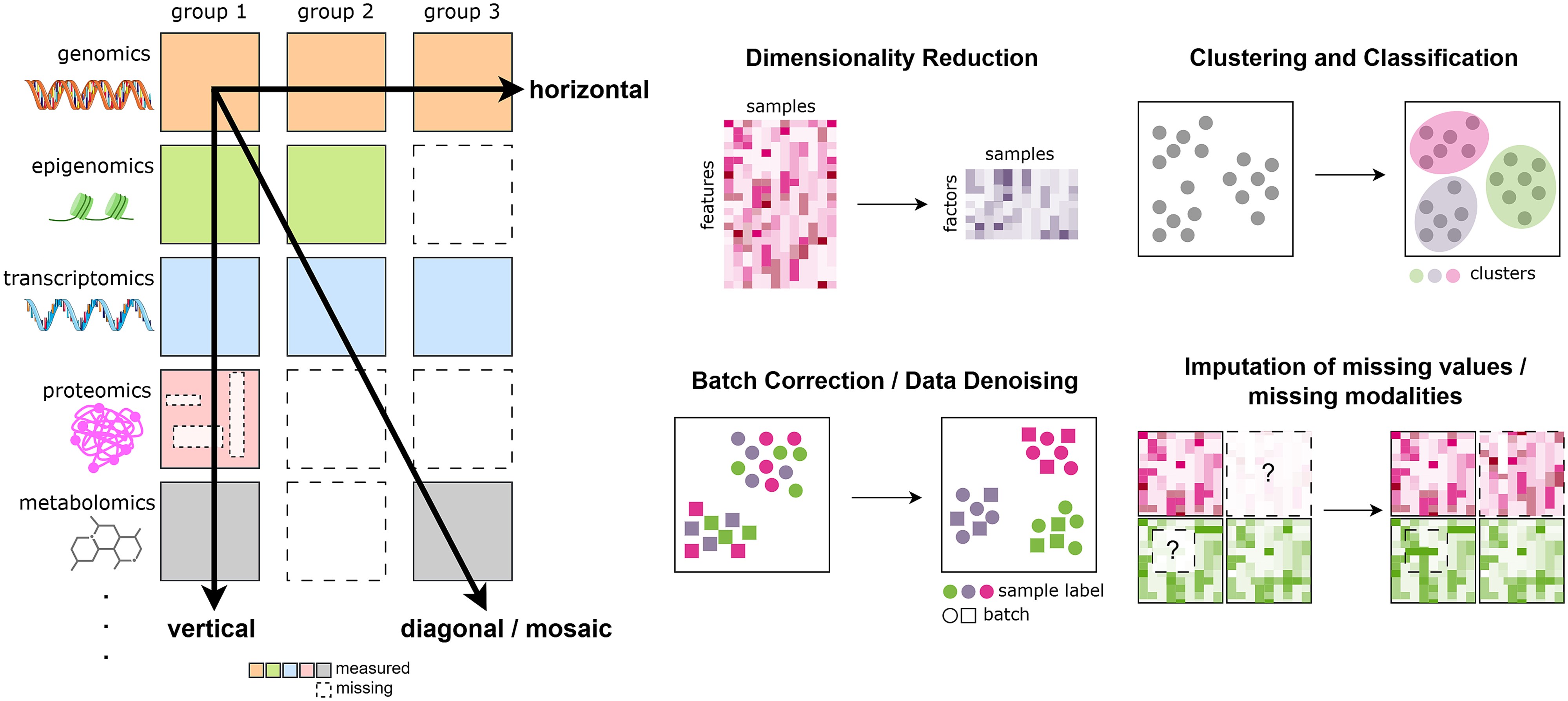
A technical review of multi-omics data integration methods from classical statistical to deep generative approaches
Multi-omics data integration review
In this review we explore cutting-edge methods for integrating complex multi-omics datasets, a key challenge...
[Read More]
New Project SYNTHESIS
Mosaic integration and synthetic generation of multi-omic data for the discovery of precision cancer medicines
Despite remarkable progress in cancer precision medicine, drug resistance and low success rates underscore the...
[Read More]
New Project SARC-RON-AI
Sarcoma Clinical Data Integration with AI-Driven Automation for the National Oncology Registry
The SARC-RON-AI project aims to develop an AI-based system to process unstructured EHRs efficiently, automating...
[Read More]
ELLIS Scholar
Emanuel accepted as an ELLIS Scholar
Emanuel has been accepted as an ELLIS Scholar with the ELLIS Lisbon Unit. He is...
[Read More]